Overview
More than 60% of human proteins are phosphorylated, and identification of the kinases and phosphatases responsible for this versatile modification has provided fundamental insights into cellular regulation. Most protein kinases phosphorylate residues that fall within a characteristic amino acid motif, which can be used to identify putative substrates. In contrast, protein phosphatases recognize their substrates with remarkable specificity, but act on phosphosites that differ widely in primary sequence. Recent studies of the PPP protein phosphatase family (which includes the major phosphatases PP1 and PP2A, as well as calcineurin) reveal that this specificity is conferred by the docking of characteristic Short Linear Motifs, or SLiMs, in substrates to conserved binding pockets on the respective phosphatase.
Calcineurin, also referred to as PPP3 or PP2B, is a metalloenzyme that utilizes a Zn2+ and an Fe2+ ion at its catalytic center to coordinate water molecules and dephosphorylate phosphoserine and phosphothreonine residues. Calcineurin is a heterodimer of catalytic (A) and regulatory (B) subunits, and is active only in the presence of Ca2+ (which binds to the B subunit) and Ca2+-bound calmodulin, which binds to an amphipathic helix in the A subunit. During activation, a conformational change occurs that relieves auto-inhibition of the active site by C-terminal sequences in calcineurin A and also exposes the LxVP SLiM binding pocket (see below).
Calcineurin acts on a select group of protein substrates by recognizing two SLiMs, PxIxIT and LxVP, which occur in disordered, accessible, protein regions, but may be hundreds of residues away from sites of dephophosphorylation and are found at variable distances and orientations with respect to each other. PxIxIT and LxVP sequences were first defined in the NFAT (nuclear factor of activated T-cells) family of transcription factors, whose nuclear accumulation is triggered by calcineurin-mediated dephosphorylation to initiate gene expression and promote T-cell activation. Subsequently, PxIxIT and LxVP motifs were identified in multiple calcineurin substrates from humans and budding yeast (S. cerevisiae), and their binding surfaces on calcineurin shown to be broadly conserved, suggesting that this characteristic mode of substrate recognition is retained throughout the fungal and animal kingdoms1.
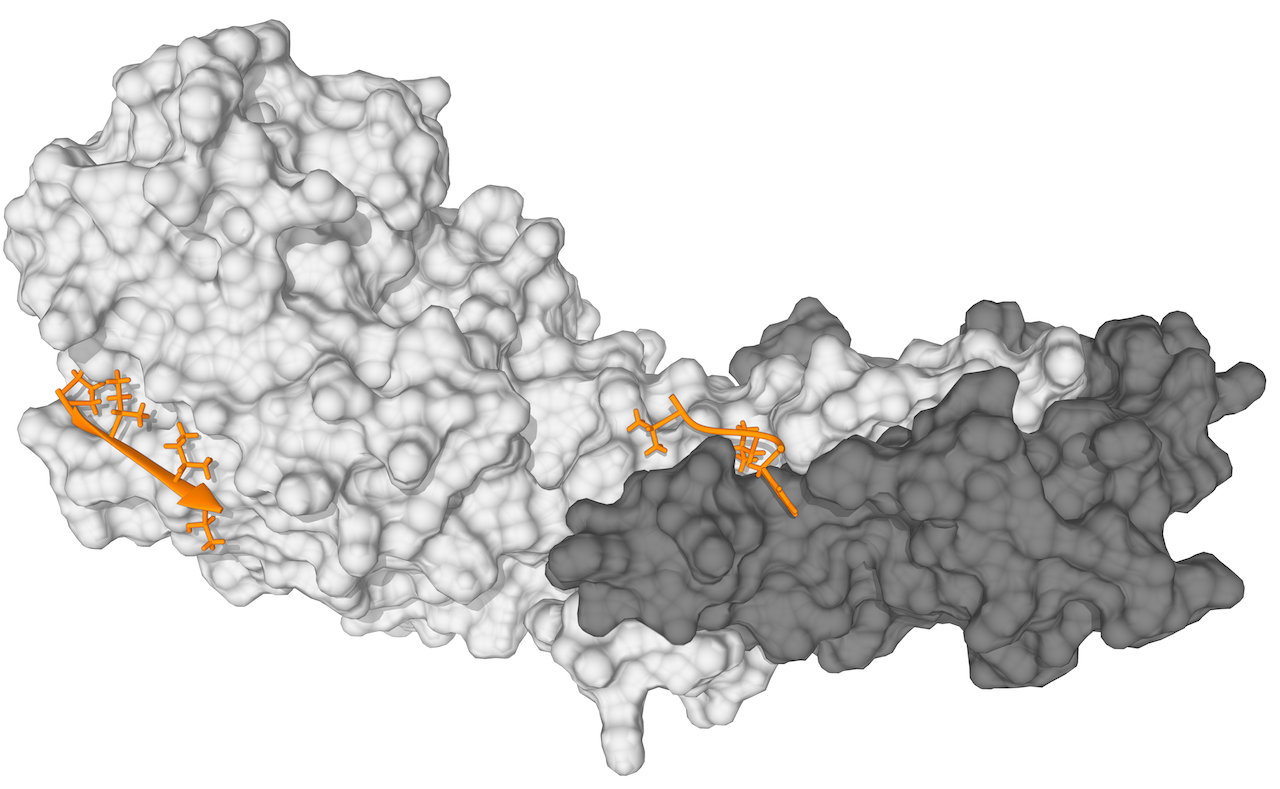 Figure 1. Structure of the PxIxIT and LxVP motif bound to calcineurin
Figure 1. Structure of the PxIxIT and LxVP motif bound to calcineurin
Crystal Structure of Calcineurin in Complex with the Calcineurin-Inhibiting Domain of the African Swine Fever Virus Protein A238L (PDB ID:4F0Z). The figure shoes the Calcineurin A (light grey) and B Subunits (dark grey) and two resolved peptides from a 43 amino acid fragment of A238L binding to the PxIxIT (orange, left) and LxVP binding pocket (orange, right).
PxIxIT sequences are found in most, but not all substrates and regulators of calcineurin. PxIxIT motifs are degenerate in sequence, but typically share a core sequence of proline (position 1), followed by two hydrophobic residues in positions 3 and 5 (I, V, L, or F), a hydrophilic residue in position 6 (T,S,H,D,E,Q,N, K or R), and a variety of amino acids in positions 2 and 4 (the “x” positions”). Only two PxIxIT sequences lacking a P in position 1 are known, in human AKAP79 and yeast RCN1. Structures of three different PxIxIT-calcineurin complexes show nearly identical interactions, with the motif forming a β strand2. However, PxIxITs vary widely in their affinity for calcineurin, with motifs from five different yeast substrates having Kds ranging from ~15-250 μM. Changing the PxIxIT affinity in Crz1, a yeast transcription factor regulated by calcineurin, alters overall calcineurin-substrate interaction strength and shifts the Ca2+ concentration-dependence of Crz1 dephosphorylation and activity in vivo3.
Recent systematic analyses of amino acid substitutions in four PxIxIT peptides determined that amino acids at the “x” positions as well as flanking residues (i.e. positions before and after the PxIxIT core) also influence affinity for calcineurin: basic residues at position 2 (K or R) increase affinity; hydrophobic or acidic residues at position -1 increase or decrease affinity, respectively; acidic or phosphorylated residues at position 9 increase affinity (Ngyuen et al submitted). A Position Specific Scoring Matrix (PSSM) generated using 33 experimentally validated human PxIxIT peptides is consistent with these findings. PxIxIT interactions are essential for dephosphorylation of most substrates: mutating the PxIxIT site in a substrate disrupts its dephosphorylation, and peptides or small molecules that compete with native PxIxIT sequences for binding to calcineurin inhibit dephosphorylation of many substrates in vitro and in vivo2,4,5.
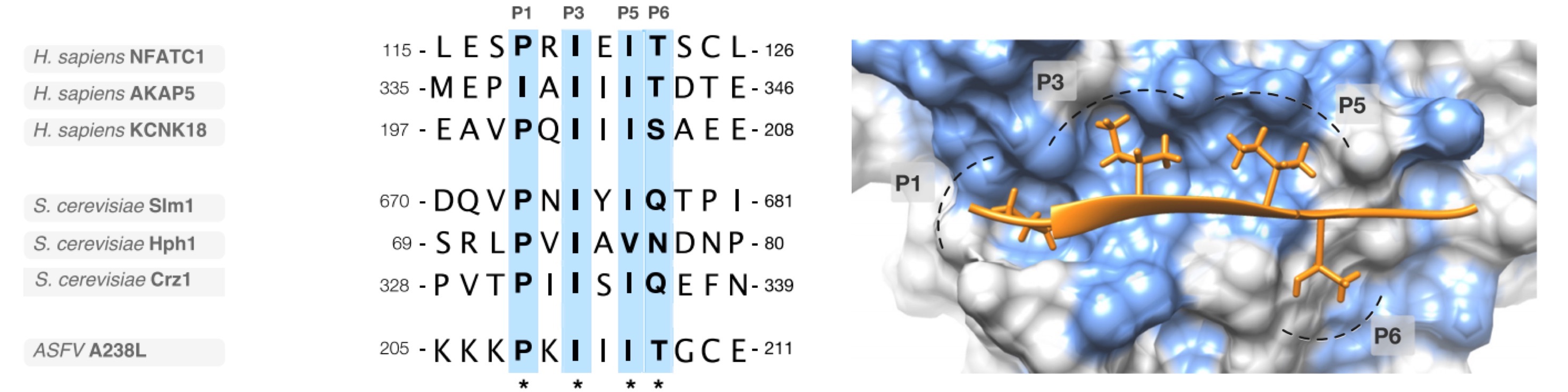 Figure 2. Structure of the PxIxIT and LxVP motif bound to calcineurin
Figure 2. Structure of the PxIxIT and LxVP motif bound to calcineurin
(A) An alignment of a representative selection of PxIxIT motif instances: Nuclear factor of activated T-cells, cytoplasmic 1 (NFATC1), A-kinase anchor protein 5 (AKAP5) and Potassium channel subfamily K member 18 (KCNK18)from human; Phosphatidylinositol 4,5-bisphosphate-binding protein SLM1 (Slm1), Protein HPH1 (Hph1) and Transcriptional regulator CRZ1 (Crz1) from yeast; and Ankyrin repeat domain-containing protein A238L from African swine fever virus (ASFV). (B) Structure of the PxIxIT binding pocket of the human calcineurin catalytic A subunit bound to the PxIxIT of African swine fever virus A238L (PDB ID:4F0Z). The peptide binds by beta-augmentation and the defined residues at P1, P3, P5 sit in a conserved hydrophobic pocket explaining the strong preferences at these positions in known PxIxIT instances (light blue surface on the domain denotes hydrophobic residues)
PxIxIT peptides form contacts along the edge of two β sheets in the calcineurin A subunit, β11 and β14. This interaction is similar to that of the RVxF SLiM with the catalytic subunit of PP1, demonstrating that these critical binding sites are evolutionarily related6. This hydrophobic docking groove is remote from the catalytic cleft, which explains the ability of PxIxIT to associate with calcineurin A in the absence of the B subunit, and to interact equally with the active and inactive forms of the enzyme. Thus, PxIxIT is critical for tethering calcineurin to substrates or regulators, such as the AKAP79 scaffold protein, but PxIxIT based-inhibitors do not interact with the catalytic center, or interfere with dephosphorylation of substrates that lack a PxIxIT site. Mutations in a conserved sequence (330NIR332 to AAA in human calcineurin A) interfere with PxIxIT binding, and severely reduce enzyme function in vivo while having no effect on catalysis3,6.
Figure 3. Sequence logo of the the PxIxIT motif
A relative binomial sequence logo constructed as log-10 of the binomial probability (probaa = binomial(k,n,p)) where k is the observed residue count at each position for a residue, n is the number of the instances of motifs and p is the backgound frequency of the residue. A positive score denotes that a residues is enriched, a negative score sugests that a residue is depleted, and score of zero suggest that a residue is occurring at the expected frequency. The logo was constructed from all experimentally characterised PxIxIT motif.
Figure 4. Frequency plot of the PxIxIT motif
A frequency plot of observed frequency of an amino acid in characterised PxIxIT motif . The plot was constructed from all experimentally characterised PxIxIT motifs. Dashed blue boxes signify significantly enriched amino acids or groupings of amino acids with shared physicochemical properties. The plot was constructed from all experimentally characterised PxIxIT motif.
The LxVP motif is shorter than PxIxIT, and is dominated by the two hydrophobic residues, L and V, which are significantly buried when bound to calcineurin. Structures of two LxVP:calcineurin complexes show this sequence binding in an extended conformation, and reveal additional hydrogen bonds formed by amino acids at the two positions (-1, -2) preceding Leucine2,7. To date, no systematic substitution analyses of the LxVP motif have been performed, however introducing tyrosine at the -1 position in Nfatc2 or DNM1L improves binding, whereas mutation of any residues in the Nfatc1 peptide (386YLAVP390) to alanine decreases binding to calcineurin8,9. A Position Specific Scoring Matrix (PSSM) generated using 34 experimentally validated human LxVP peptides is consistent with an expansion of the motif to 6 or 7 positions and shows a preference for hydrophobic/polar residues at the -1 and -2 positions. Two auto-inhibitory sequences in calcineurin A bind to the LxVP binding pocket in calcineurin: 1) Structural analysis of the β isoform of human calcineurin A shows that a C-terminal peptide, 416ARVFSVLR423 fills the LxVP-binding pocket with FSVL overlaying the LxVP in previous structures10. This auto-inhibitory sequence, or AIS, blocks the LxVP pocket under basal conditions and helps maintain calcineurin in the inactive state. 2) A unique isoform of human calcineurin Aβ, CnAβ1, which is generated via alternative 3’ end mRNA processing, contains a C-terminal 460MQLAVP465 sequence that auto-inhibits this isoform by directly blocking the LxVP-binding pocket11.
The LxVP motif binding pocketLxVP peptides bind to a hydrophobic channel formed at the intersection of the calcineurin A and B subunits. Under basal conditions, the LxVP binding groove is occluded by the AIS, and becomes available for substrate binding only after enzyme activation with Ca2+ and Ca2+/calmodulin. Structural and biochemical analyses of LxVP-calcineurin interactions show that immunosuppressants, Cyclosporin A and FK506, in complex with their respective small binding protein (immunophilin), block the LxVP-binding pocket of calcineurin2. Residues 352W and 356F in human calcineurin Aα, which occur in the B-subunit binding region of calcineurin A, help to form hydrophobic pockets for the L and V residues2. Substitution of alanine for either of these residues reduces binding to both LxVP peptides and immunosuppressant-immunophilin complexes, and compromises in vivo function of the enzyme, while also providing some resistance to immunosuppressants8. Because Cyclosporin A and FK506 inhibit calcineurin from multiple species, and abrogate all known calcineurin-mediated dephosphorylation events, engagement of the LxVP-binding pocket by substrates is thought to be essential for dephosphorylation.
Figure 5. Sequence logo of the the LxVP motif
A relative binomial sequence logo constructed as log-10 of the binomial probability (probaa = binomial(k,n,p)) where k is the observed residue count at each position for a residue, n is the number of the instances of motifs and p is the backgound frequency of the residue. A positive score denotes that a residues is enriched, a negative score sugests that a residue is depleted, and score of zero suggest that a residue is occurring at the expected frequency. The logo was constructed from all experimentally characterised LxVP motif.
Figure 6. Frequency plot of the LxVP motif
A frequency plot of observed frequency of an amino acid in characterised LxVP motif. The plot was constructed from all experimentally characterised LxVP motifs. Dashed blue boxes signify significantly enriched amino acids or groupings of amino acids with shared physicochemical properties. The plot was constructed from all experimentally characterised LxVP motif.
Several experimentally identified calcineurin-binding peptides contain overlapping LxVP and PxIxIT motifs, i.e. LxVPxIxIT, or combination motifs. One such motif from CACNA1H, 2260HLTVPSFAFEPLD2272, was demonstrated to have both LxVP and PxIxIT-type binding characteristics to calcineurin in vitro. Because of the large distance between the LxVP and PxIxIT binding pockets on calcineurin, these peptides are proposed to bind either in the PxIxIT or LxVP mode, rather than binding simultaneously to both sites.
Coordination of PxIxIT and LxVP motifsAs first shown for NFAT family members, PxIxIT and LxVP motifs both contribute to protein recognition and dephosphorylation by calcineurin. While no calcineurin:substrate complex has yet been solved, a structure of calcineurin bound to a 39 amino acid region of A238L, a protein produced by African Swine Fever Virus that inhibits the phosphatase, has been analyzed2. The A238L peptide binds tightly to calcineurin (Kd=4nM) via a PxIxIT (206PKIIIT211) and an LxVP (228FLCVK232) motif, with each SLiM contributing approximately equally to overall affinity and inhibition of calcineurin in vivo. Thus, stable binding to calcineurin at one or both SLiM docking sites prevents dephosphorylation of native substrates, which interact only transiently with the enzyme. Furthermore, proteins with one ‘consensus’ SLiM of relatively high affinity may require only a poorly matching second SLiM for recognition and dephosphorylation by calcineurin.
PxIxIT and LxVP motifs play distinct roles during dephosphorylation. By binding under both basal and signaling conditions, PxIxIT motifs target calcineurin to substrates/regulators or to protein complexes that contain substrates. For example, the human scaffold protein, AKAP79, co-binds calcineurin and two substrates (the L-type Ca2+ channel and PKA RII regulatory subunit) that lack PxIxITs but contain LxVP motifs. In contrast, LxVP motifs, which can only bind active calcineurin, may help orient the phosphosite towards the catalytic center of calcineurin for dephosphorylation. This mechanism is supported by computational modelling of calcineurin in complex with a substrate (the LxVP-containing peptide derived from the PKA RII regulatory subunit), and predicts that dephosphorylated residues are 9-15 residues C-terminal to the LxVP motif2. However, only a subset of substrates fit this paradigm (PKA RII, RCAN1-3). In NFAT, for example, the PxIxIT is located 200 amino acids N-terminal to the LxVP, with 13 calcineurin-dependent phosphorylation sites in between12. Thus, other modes of substrate engagement by calcineurin likely await discovery. References
1 "Cracking the phosphatase code: docking interactions determine substrate specificity." Roy J, Cyert MS. Science signaling. (2009) 2(100):re9.
2 "The molecular mechanism of substrate engagement and immunosuppressant inhibition of calcineurin." Grigoriu S, Bond R, Cossio P, Chen JA, Ly N, Hummer G, et al. PLoS Biol. (2013) 11(2):e1001492.
3 "A conserved docking site modulates substrate affinity for calcineurin, signaling output, and in vivo function." Roy J, Li H, Hogan PG, Cyert MS. Molecular cell. (2007) 25(6):889-901.
4 "Affinity-driven peptide selection of an NFAT inhibitor more selective than cyclosporin A." Aramburu J, Yaffe MB, Lopez-Rodriguez C, Cantley LC, Hogan PG, Rao A. Science. (1999) 285:2129-33.
5 "Identification of small-molecule inhibitors of calcineurin-NFATc signaling that mimic the PxIxIT motif of calcineurin binding partners." Matsoukas MT, Aranguren-Ibanez A, Lozano T, Nunes V, Lasarte JJ, Pardo L, et al. Science signaling. (2015) 8(382):ra63.
6 "Structural delineation of the calcineurin-NFAT interaction and its parallels to PP1 targeting interactions." Li H, Rao A, Hogan PG. Journal of molecular biology. (2004) 342(5):1659-74.
7 "Investigating the human Calcineurin Interaction Network using the piLxVP SLiM. Scientific reports." Sheftic SR, Page R, Peti W. Sci Rep. (2016) 6:38920.
8 "A conserved docking surface on calcineurin mediates interaction with substrates and immunosuppressants." Rodriguez A, Roy J, Martinez-Martinez S, Lopez-Maderuelo MD, Nino-Moreno P, Orti L, et al. Molecular cell. (2009) 33(5):616-26.
9 "A calcineurin docking motif (LXVP) in dynamin-related protein 1 contributes to mitochondrial fragmentation and ischemic neuronal injury." Slupe AM, Merrill RA, Flippo KH, Lobas MA, Houtman JC, Strack S. The Journal of biological chemistry. (2013) 288(17):12353-65.
10 "Cooperative autoinhibition and multi-level activation mechanisms of calcineurin. Li SJ, Wang J, Ma L, Lu C, Wang J, Wu JW, et al. Cell research. (2016) 26(3):336-49.
11 "The unique C-terminus of calcineurin isoform CNAbeta1 confers non-canonical regulation of enzyme activity by Ca2+ and calmodulin." Bond R, Ly N, Cyert MS. The Journal of biological chemistry. (2017) 292(40):16709-16721.
12 "Concerted dephosphorylation of the transcription factor NFAT1 induces a conformational switch that regulates transcriptional activity." Okamura H, Aramburu J, Garcia-Rodriguez C, Viola JP, Raghavan A, Tahiliani M, et al. Molecular cell. (2000) 6(3):539-50.
