Overview
Eukaryotic cell cycle progression is largely regulated by the two oscillating systems – the Cyclin-Cdk phosphorylation system and the anaphase-promoting complex/cyclosome (APC/C) ubiquitination system. The APC/C is a large multi-subunit E3 ubiquitin ligases that poly-ubiquitinate substrates, targeting them to the proteasome thereby promoting their degradation. All fully sequenced eukaryotic genomes contain at least two APC/C substrate recognition modules known as co-activators (though some proteomes have more). These two ever-present co-activator protein, known as Cdc20 and Cdh1, recognise substrates, recruit them to the APC/C and position them to allow their efficient poly-ubiquitination by APC/C-bound E2 ubiquitin ligases. The APC/C is not constitutively active and numerous regulator mechanisms co-operate to limit activity to a window of time between pro-metaphase and the end of G1. During pro-metaphase and metaphase, Cdc20 acts as the APC/C activator, changes in the phosphorylation state of the APC/C and Cdh1 results in a activator change from Cdc20 to Cdh1 from late metaphase onwards.
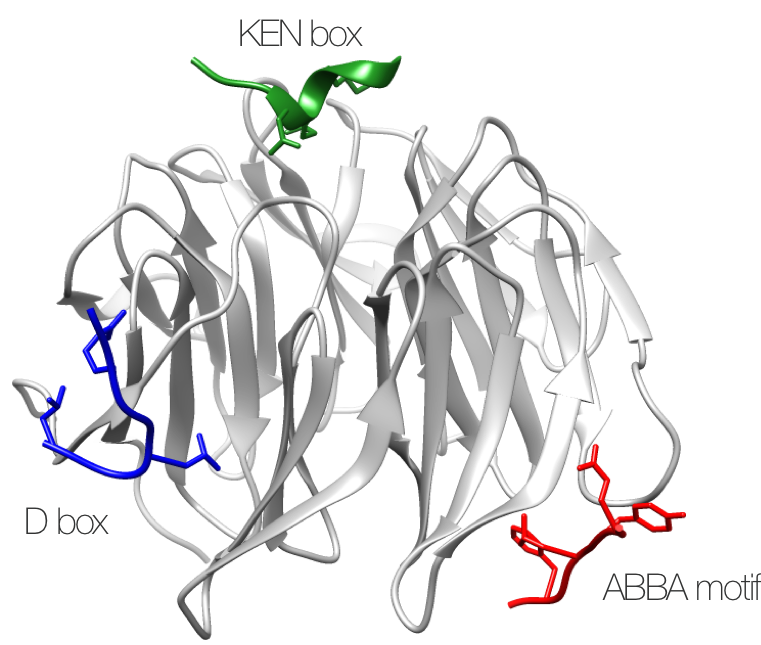
Figure 1. Structure of the three APC/C degrons
Structure of S. cerevisiae APC/C activator protein CDH1 (Cdh1) bound to the three degrons of APC/C-CDH1 modulator 1 (Acm1) (PDB:4BH6).
The APC/C activators recognise short peptides in the disordered regions of their substrates, a subclass of short, linear motifs (SLiMs) known as degrons (derived from degradation motif). These APC/C degrons acts as specific signals to the cell that a given protein should be destroyed in a cell cycle-dependent manner. The APC/C activators contain three known short, linear motif (SLiM) binding pockets. The degrons that bind these pockets are known as the D box, KEN box and ABBA motif. Each degron binds to a distinct binding sites on the surface of the WD40 repeat domain of the activator (Figure 1). D box and KEN box motifs are recognised by all tested activators. However, evolutionary evidence suggests that there exists activators that do not bind to specific degrons (Figure 2). The metazoan Cdh1 appears to have lost its ABBA motif binding pocket and none of the known ABBA motifs tested could bind Cdh1. Similarly, the third meiosis specific activator of yeast AMA1 is missing key ABBA motif binding residues in the ABBA binding pocket. The Cdc20 orthologue (FZY-1) in C. elegans also appears to have lost its KEN binding pocket however this has yet to be investigated. Human and closely related metazoan species also have a Cdc20 paralogue (known as CDC20B) that appears to have lost its APC/C degron binding ability (and the motifs required for APC/C binding such as the C box and the IR tail). The function of these paralogues are poorly understood.
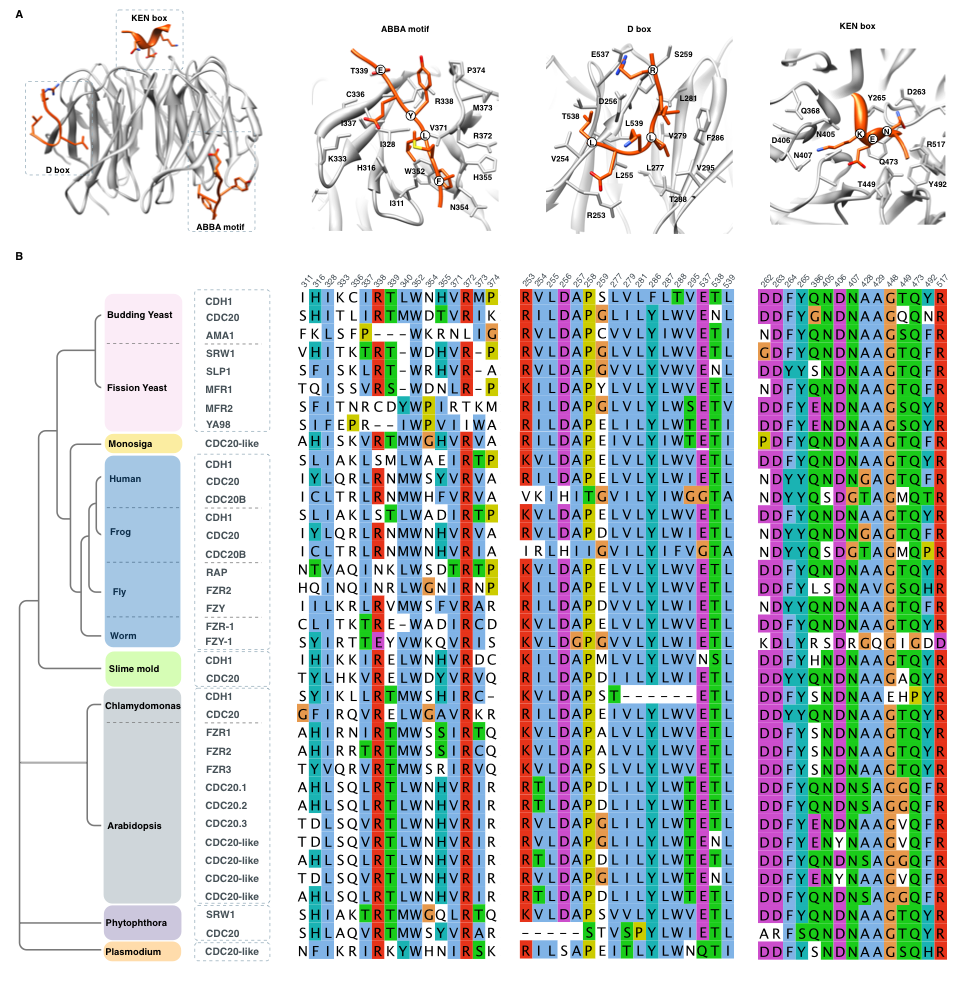
Figure 2. Conservation of the three APC/C degron binding pockets
(A) Structure of an activator bound to the three APC/C degrons showing numbering of the residues in the binding pocket. Residues in the binding pocket were calculated from the structure of the APC/C activator protein CDH1 (Cdh1) bound to APC/C-CDH1 modulator 1 (Acm1) (PDB:4BH6) and consequently the numbering corresponds to the yeast APC/C activator protein Cdh1. (B) Conservation of the residues calculated to be in contact with the three APC/C degrons across several eukaryotic proteomes.
Members of multi member motif-binding domain families such as the WD40 repeats often recognize a common core motif (e.g. SH2 domains recognise phosphotyrosine, SH3 domain recognise polyproline PxxP motifs, and PDZ domains recognise hydrophobic C-terminal motifs). However, different domain family members can bind distinct subsets of peptides despite having the same core-binding pocket. The core-binding pocket is complementary to the required position in the motif, shared across all peptides binding to the domain family, and interactions with this pocket contributes the majority of the specificity and affinity of binding. Evolutionary refinement of the binding surfaces of these domains out side the core pocket contribute the subtle specificity determinants of the interaction discriminating between various motif-containing peptides to allow complete separation, partial overlap, or complete overlap of the targets of homologous domains. For example, despite complete conservation of the D-box binding pocket across all Cdc20-like proteins there exist instance of D-box peptides, such as the N-terminal D-box of Acm1, that are specific for either Cdc20 or Cdh1. However, this not fully understood for most APC/C degrons with the exception of the ABBA motif.
D box
The D box was first characterised in Cyclin B by Glotzer et al. The D box degrons bind to a pocket situated between blades 1 and 7 of the WD40 repeat of the activator (Figure 1 - blue peptide). The binding pocket contains two important regions - an acidic and a hydrophobic patch that are occupied by the the basic and aliphatic residues of the classical D box consensus. The large majority of characterised D boxes follow the consensus RxxLxx[ILV]. There are strong preferences outside of the consensus (Figure 3). The long recognised +9 asparagine occurs in ~30% of residues and should not be considered part of the consensus. Almost half of all D boxes have a +3 position alanine or a proline. The +6 position has a strong preference for an acidic residues. There is a preference for a threonine or a serine residue in the +8 position. There is an increased frequency of basic residues in the N-terminal region and of asparagine residues in the C-terminal region of the D box containing peptides. The optimal D box would follow a consensus R[KR][AP]Lx[DE][ILV][TS]N. Such a peptide would likely have a relatively high binding affinity. Phosphorylation of the +2 position has been shown to inhibit activator binding, and conversely, phosphorylation of the +6 position has been shown to promote activator binding.
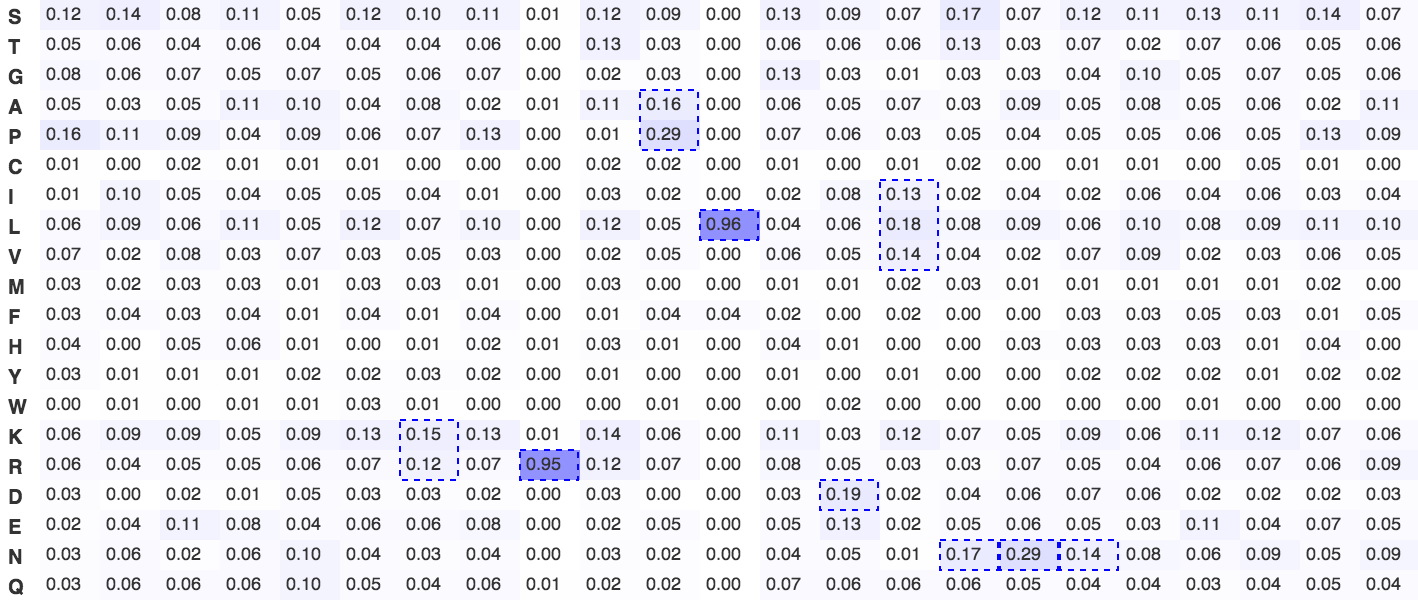
Figure 3. Frequency plot of the D box degron
A frequency plot of observed frequency of an amino acid in characterised d boxes. The plot was constructed from all experimentally characterised D box degrons. The strongly favoured arginine and leucine of the over-predictive RxxL consensus are clearly visible. Dashed blue boxes signify significantly enriched amino acids or groupings of amino acids with shared physicochemical properties.
In recent years, numerous non-canonical D box instances have been characterised suggesting that the lack of a consensus residue in a certain position can be compenstated for in other non-consensus positions. It appears that there exists a set of resides at each position that are allowed. Some residues contribute more than other to the binding, so a functional D box can have weak or strong residues at each position. The better the residue that is present at each position the better the binder. When a peptide has a strong residue at a given position, the peptide can tolerate a weaker residue at another position, as it compensates for it. The Hsl1 super D box has the optimal residues in nearly all positions and is the strongest known binder. The Cyclin A D box is not transplantable and only works as a part of an avidity driven interaction. It only has the bare consensus. Whereas the homologous Cyclin B D box which is transplantable and functional has optimal residues in many positions. Most non-canonical D boxes don't fit the consensus but compensate by having strongly preferred residues in multiple other non-classical positions.
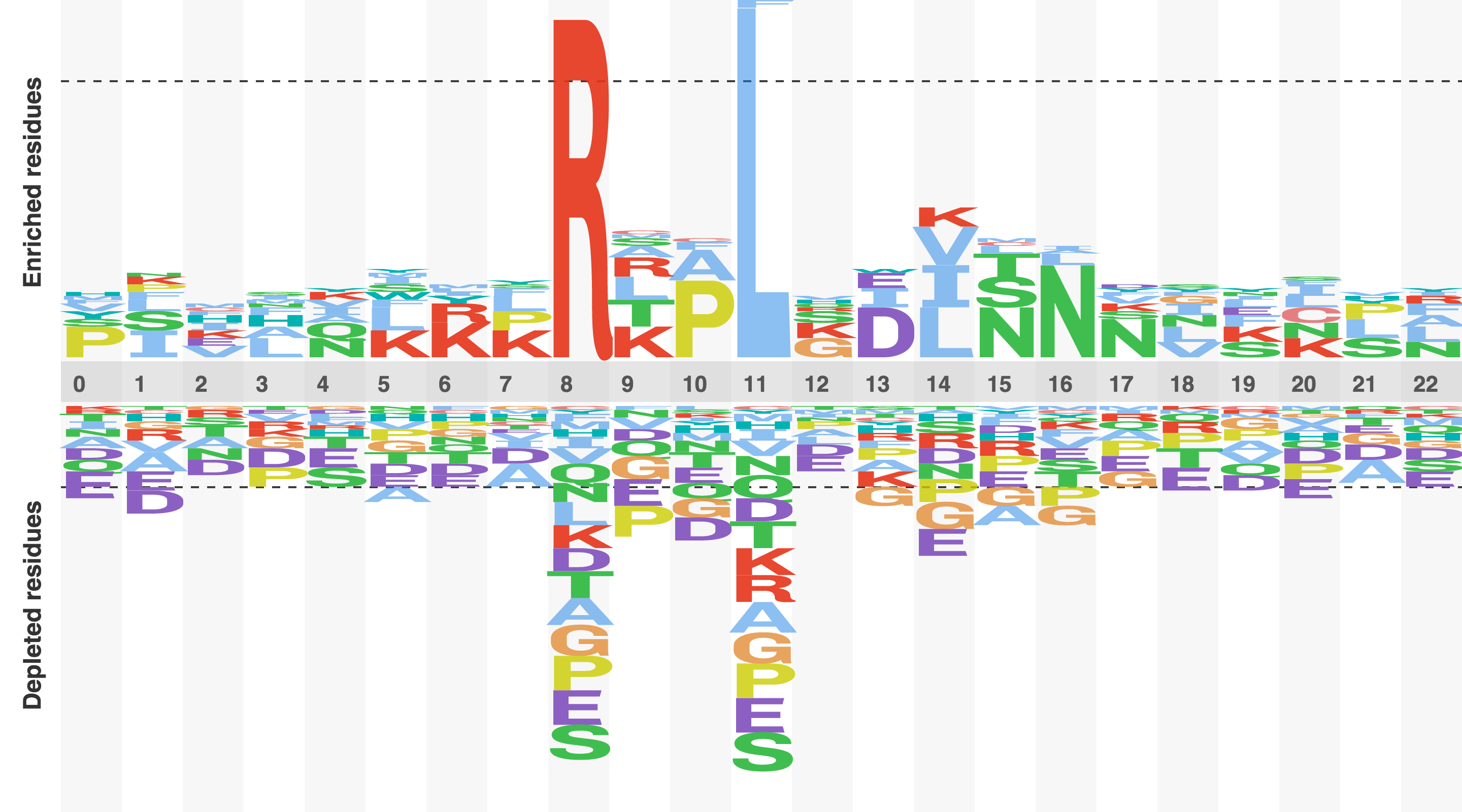
Figure 4. Sequence logo of the D box degron
A relative frequency sequence logo constructed by subtracting the expected frequency of an amino acid from the observed frequency of an amino acid. A positive score suggests that a residues is enriched , a negative score sugests that a residue is depleted, and score of zero suggest that a residue is occurring at the expected frequency. The logo was constructed from all experimentally characterised D box degrons.
KEN box
A decade after the discovery of the D box Pfleger et al characterised the KEN box. The KEN box binds in a depression at the centre of the top surface of the wheel-like WD40 repeat domain. The motif is very strictly defined for a short, linear motif and rarely deviates from the classical KEN consensus, particularly in the glutamic acid and asparagine positions, which are present in all characterised KEN box motif instances. The lysine position has been observed to allow other residues - usually aspartic acid, glutamine or asparagine (which has been crystallised in one of the dimers of the Acm1-Cdh1 tetrameric structure). Furthermore, a -1 position glycine, GxEN motif has been shown to compensate for the lack of an optimal residue in the lysine position. The flanking regions of the motif show statistically significant enrichment in several positions. Over half of all characterised KEN boxes have an acidic residues (aspartic acid, glutamic acid and asparagine) in the -1 position. The +4 position is enriched for hydrophobic residues with ~40% of instances conforming to this preference. Finally, the +6 position often contains a proline residue. Phosphorylation of the +5 position has been hypothesised to stabilise the bound conformation of the KEN box and may be a regulatory position for the motif.
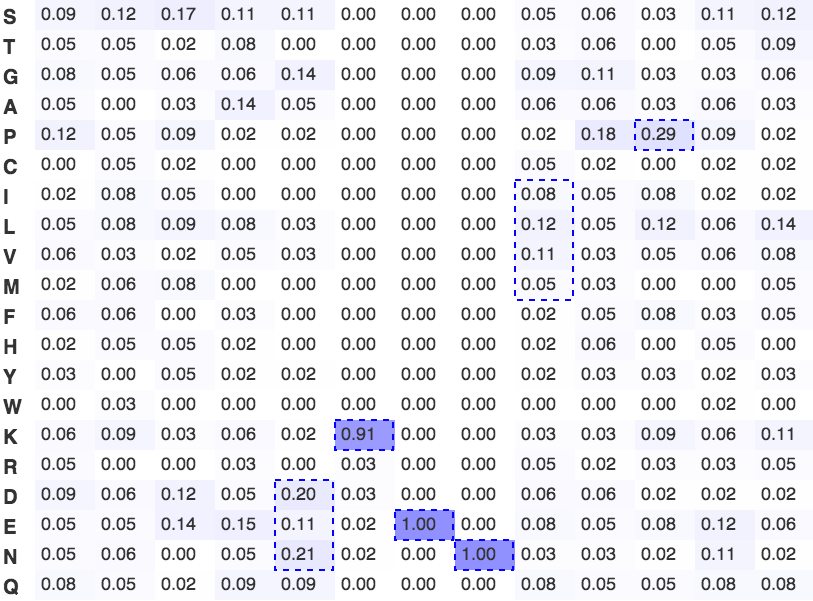
Figure 5. Frequency plot of the KEN box degron
A frequency plot of observed frequency of an amino acid in characterised KEN boxes. The plot was constructed from all experimentally characterised KEN box degrons. The strongly favoured lysine, glutamic acid and asparagine of the KEN consensus are clearly visible. Dashed blue boxes signify significantly enriched amino acids or groupings of amino acids with shared physicochemical properties.
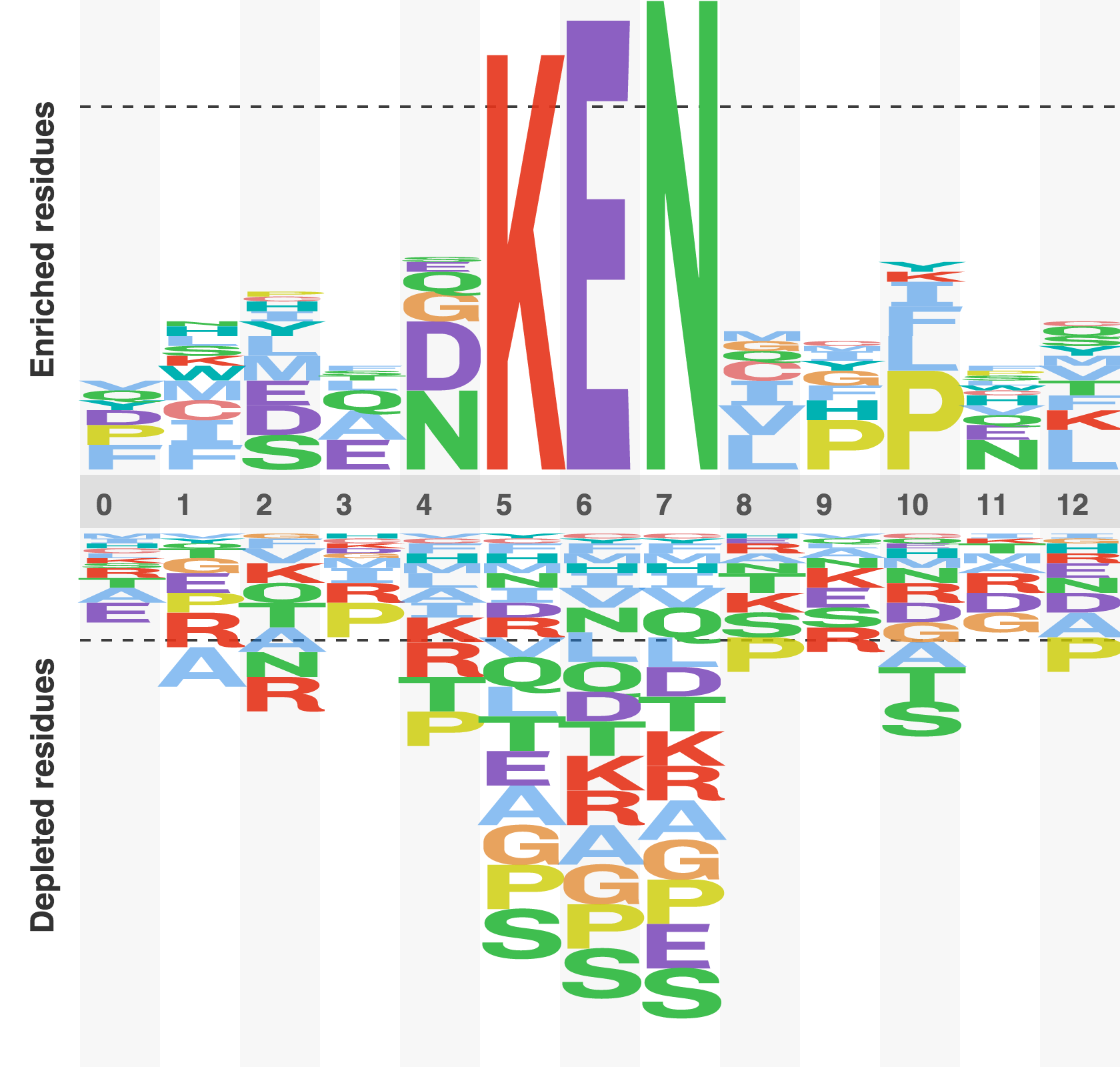
Figure 6. Sequence logo of the KEN box degron
A relative frequency sequence logo constructed by subtracting the expected frequency of an amino acid from the observed frequency of an amino acid. A positive score suggests that a residues is enriched , a negative score sugests that a residue is depleted, and score of zero suggest that a residue is occurring at the expected frequency. The logo was constructed from all experimentally characterised KEN box degrons.
See a list of the experimentally characterised KEN boxes here.
ABBA motif
The ABBA motif was the most recently characterised APC/C degron. The full characterisation of the binding preference of the pocket is limited by the small number of known ABBA instances, however, it appears that the core residues of the ABBA peptide are 3 evolutionarily constrained positions (the [ILMVP][FHY]x[DE] positions shared by Clb5, Cyclin A, BubR1, Bub1 and Acm1). The pocket in contact with these residues is largely conserved. However, a single residue in the flanking regions of the ABBA motif core has the potential to switch the specificity of a peptide from a Cdc20 binder to a Cdh1 binder. The Cdc20 of metazoan lineage and Cdh1 of the fungal lineage has a additional preference for an aromatic or aliphatic residue in to the N-terminal region of the motif expanding the consensus to [FILV]x[ILMVP][FHY]x[DE]. Remarkably, the specificity of the Cdc20 of the fungal lineage has diverged resulting in a preference for peptides matching [KR]xx[ILV][FHY]x[DE]. Observations in Yeast offers an insight into these diverging specificities of APC activators. Firstly, modeling based on the Acm1-Cdh1 complex offers an explanation for the specificity of the Clb5 ABBA motif for Cdc20 over the Cdh1 target Acm1. Several mutations in a loop at the back of the ABBA binding pocket of yeast Cdc20 has created an acidic binding surface that is not present in Yeast Cdh1 (Figure 7). Clb5, a yeast Cdc20 target has a complementary basic patch that is absent in Acm1, suggesting that this difference between Cdh1 and Cdc20 may promote a specific Cdc20-Clb5 interaction. Indeed, replacement of the acidic loop in Cdc20 with the Cdh1 residues inhibits Clb5 binding to Cdc20. Consequently, yeast Cdc20 appears to have a preference for ABBA motifs with a lysine at the -1 position and lacks the preference for a hydrophobic residue at position 1.
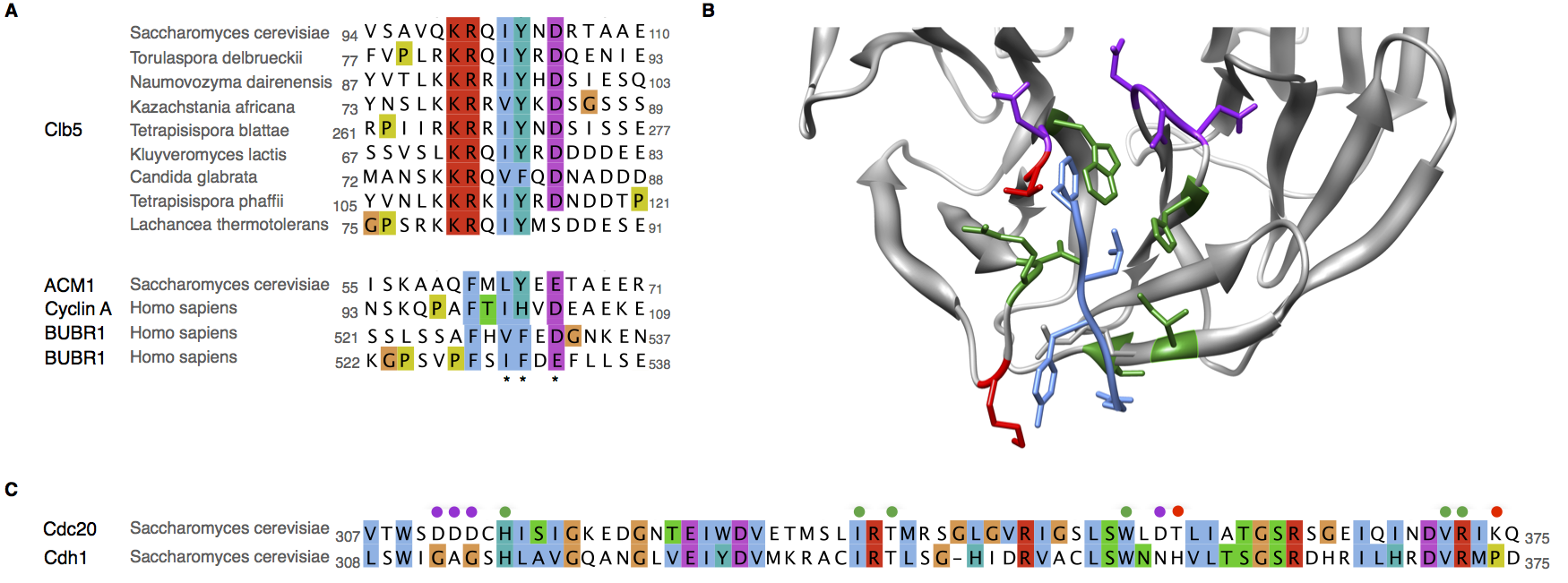
Figure 7. Change in specificity of the ABBA motif binding pocket
(A) Alignment of the experimentally characterised ABBA motif degrons showing the preferred residues in the yeast Cdc20 targets and the human Cdc20 targets. (B) Structure of the ABBA motif binding pocket showing the shared (green), different (red) and specificity determining (purple) ABBA motif-contacting residues between yeast Cdc20 targets and the human Cdc20. (C) Alignment of the ABBA motif binding pocket region of the WD40 repeat showing the shared (green), different (red) and specificity determining (purple) ABBA motif-contacting residues between yeast Cdc20 targets and the human Cdc20.
In human, the ABBA motifs of Cyclin A, Bub1 and BubR1 do not bind to Cdh1 and no ABBA instances for any metazoan Cdh1 have been characterized. Several features of the binding pocket of human Cdh1 are conserved (Figure 2), however, a lysine in the floor of the ABBA binding pocket conserved across all characterized ABBA binding domains (human Cdc20, yeast Cdh1 and yeast Cdc20) is a serine in human Cdh1 and the portion of blade contacting the conserved lysine, conserved in human Cdc20, yeast Cdh1 and yeast Cdc20, has diverged significantly in Cdh1. This region forms part of the contact for position 4 of the ABBA motif (Interestingly, this region is also disrupted in yeast meiotic activator Ama1). In the absence of a known metazoan Cdh1 binding ABBA motifs or structural characterization of the WD40 repeat of human Cdh1 it is difficult confidently predict the effect of such mutations on the specificity of the pocket. However, it does raises that possibility that human Cdh1 can no longer act as an ABBA binding module or that the specificity of the human Cdh1 ABBA binding pocket has diverged significantly.
See a list of the experimentally characterised ABBA motifs here.
Other motifs
Several other pockets on the APC/C complex surface outside of the activator are also motif-binding sites, for example, the LRRL site, IR tail site and KILR motif. However, these sites are largely regulatory in nature and bound by the activators, E2 ubiquitin ligases, inhibitors and accessory proteins of the APC/C. These binding pockets, are utilised by substrates, for example, IR tails are used by Serine/threonine-protein kinase Nek2 (NEK2) and Kinesin-like protein KIF18A (KIF18A) [23288039], though they appear to be exceptions. Other potential binding motifs observed to modulate APC/C-substrate interactions such as the TEK box and the IM remain to be fully characterised.
See a list of the experimentally characterised non-canonical APC/C degrons here.
